 
In case you want to process more than one molecule (e.g. If the RDkit molecule object contains more than one molecule, you won't be able to build the ZMatrix. Peptides are still ok, but computing proteins might take a while. The algorithm is quite inefficient for large molecules. 
Requirementsĭifferent examples for how to use this python class can be found in examples. The calculation of all these virtual coordinates can be carried out with this python class. By convention, the Cartesian coordinates of our molecule A-B-C-D-E will be shifted to the position of Z. Second, we need the torsion angles X-Y-Z-A, Y-Z-A-B, Z-A-B-C, the angles Y-Z-A, Z-A-B and the bond length Z-A. First, we need 3 Cartesian coordinates X, Y, and Z in the reference frame, which can be basically anything in the lab coordinate system, as long as it is well defined. a ligand) with 5 atoms: A-B-C-D-E and we want to convert them back to Cartesian space. For instance, let's say we have the Zmatrix coordinates for a molecule (e.g. These coordinates are also called 'virtual coordinates'. coordinates in a host or protein molecule) and 3 torsion angles, 2 angles and 1 bond length with respect to these coordinates. The algorithm is described in detail in these two articles and Ī general remark on transformations is that the conversion from ZMatrix coordinates to Cartesian coordinates requires 3 reference Cartesian coordinates (e.g. In order to circumvent this, I implemented the Natural Extension Reference Frame algorithm, which minimizes these errors. One difficulty in this context is to avoid numerical instabilities (for instance through round-offs), since these might propagate through the Z-Matrix during the conversion. However, the back conversion is more tricky and somewhat error prone. The transformation from Cartesian coordinates to ZMatrix coordinates is straightforward. Therefore, the openmm package is used for dealing with units and enforcing the correct ones during every step. If one has Cartesian coordinates for the RDKit molecule, one can also generate the Z-Matrix coordinates or convert the Z-Matrix coordinates back to Cartesian coordinates.ĭuring all conversions, one easily can make a mistake by using wrong units. The bond vector definition is here: def get_bond_vector(r, a, t):Īgain, the only part I don't understand is # get local axis system from 3 coordinates.With this little python implementation, RDkit molecules can be converted to a Z-Matrix topology representation. Here is some context of how this function is used: bond_vector = get_bond_vector(atom.rval, atom.aval, al)ĭisp_vector = np.array(np.dot(bond_vector, _axes))Ītom.coords = + disp_vector What is "getting local axis system from 3 coordinates"? U23c21 = get_ucp(u23, u21) # unit cross product Print('\nError: Co-linear atoms in an internal coordinate definition') U23 = get_u12(coords2, coords3) #calculating vector between that points 2-3 U21 = get_u12(coords1, coords2) #calculating vector between that points 1-2 However, in his work, there is a mathematical part that I don't understand: # get local axis system from 3 coordinatesĭef get_local_axes(coords1, coords2, coords3): While searching I found TMPChem's work on GitHub and it does exactly what I want. If there aren't any mistakes up to this point, how I will calculate Cartesian coordinates of the 4th atom using these $x,y,z$ values? To calculate $x,y,z$ values from the formulas x = r * sin(theta) * cos(phi) I am not sure if I should calculate this as z2 + s(angle) or z2 - s(angle) and what it depends on, if both are possible.įor the 4th atom, I use spherical coordinates r, theta, phi = (0.976, 96.572, -179.995) The 3rd atom must have coordinates that are something like this if read correctly: 3 H 0 distance*sin(angle) z2+distance*cos(angle) How should I treat the 3rd and 4th atoms? My question is after setting first atom as 0,0,0 1 O 0 0 0Īnd the second one as 0,0,(distance from first) to put it on the z-axis 2 O 0 0 1.45335189476 This is converting a Z-matrix to Cartesian coordinates. Now I need to perform the reverse operation and use this Z-matrix as input and define $x,y,z$ coordinates for each atom. Then, the script constructs a Z-matrix with them, like this: Z-mat : 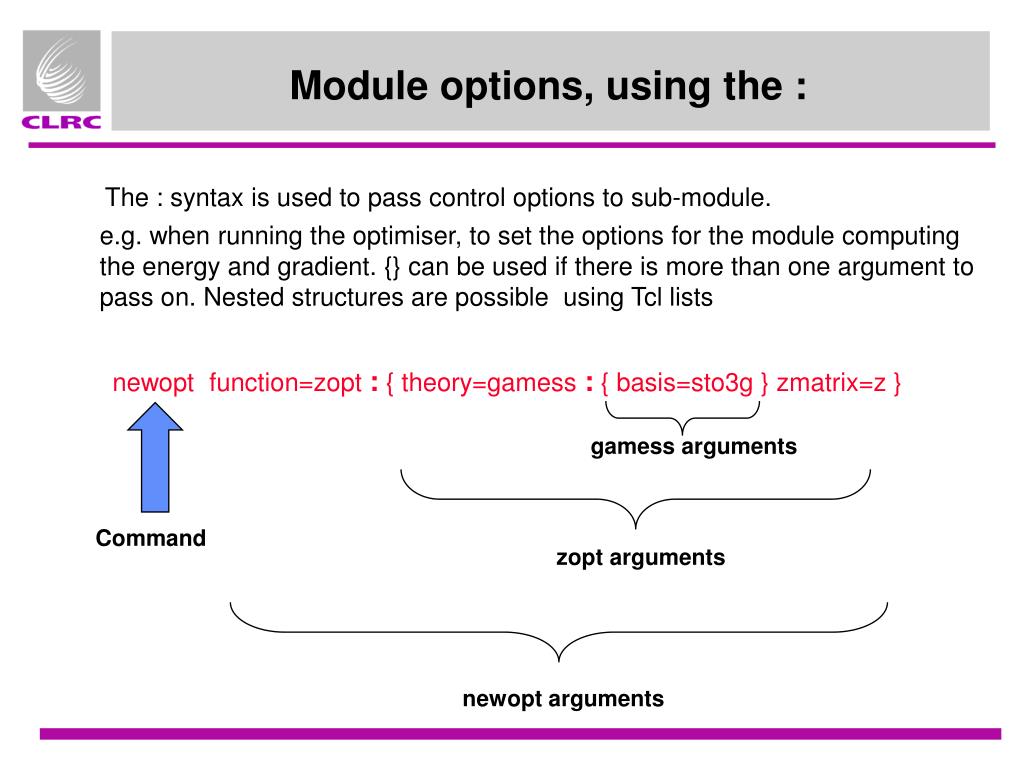
These are $x,y,z$ coordinates of H2O2 molecule. To begin with, I wrote a script that gets Cartesian coordinates of molecule as input in the below. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
